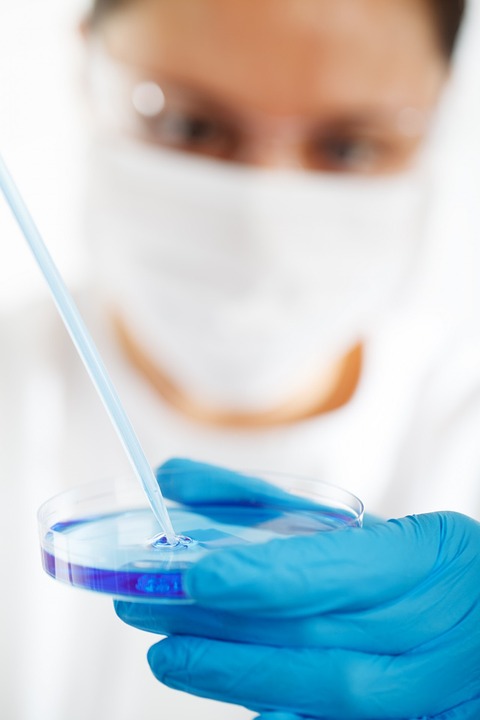
Schmid College of Science and Technology
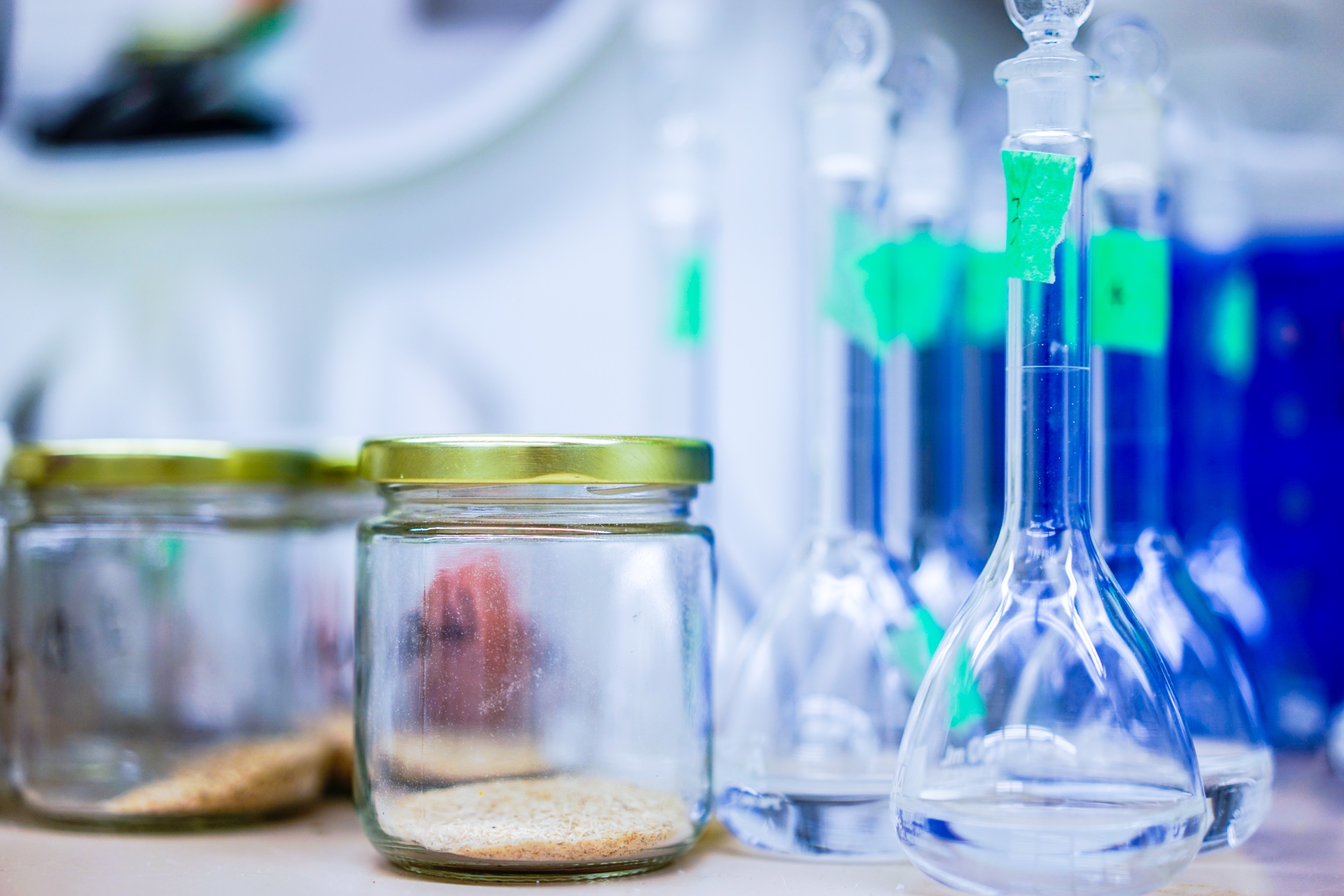




chapman

Computational Cancer Biology
Protein Kinases and Cancer Research
Molecular Chaperones and Cancer Research
Translational Bioinformatics and Drug Discovery
Exploring the molecular and cellular basis of metastasis
Chapman University

Schmid College of Science and Technology
Chapman University

Our research efforts are in the areas of Protein Biophysics, Computational Cancer Biology, Translational Bioinformatics, Computational Medicine and Drug Discovery. We focus on the development and application of computational biology approaches for (a) system-based analysis of evolutionary, genetic, and molecular signatures of human disease; (b) design and discovery of targeted and personalized cancer therapeutics; (c) integration of computational biology and chemical biology approaches in translational cancer research.
Approach & Technology
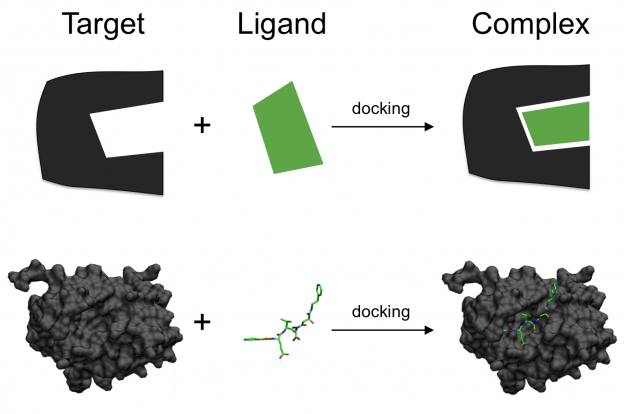
Translational Bioinformatics and Drug Discovery
Posted: September 14, 2017
The greatest challenge of the postgenomic era is to understand the function of genes and gene products in multiple organisms –including humans– both from fundamental and applied perspectives. This will […]
Research Highlights
Computational Cancer Biology
Posted: September 14, 2017
The challenge of understanding biological systems at the molecular and systems level as well as the integration of computational and experimental approaches for bridging basic and clinical cancer research is […]
Blogs
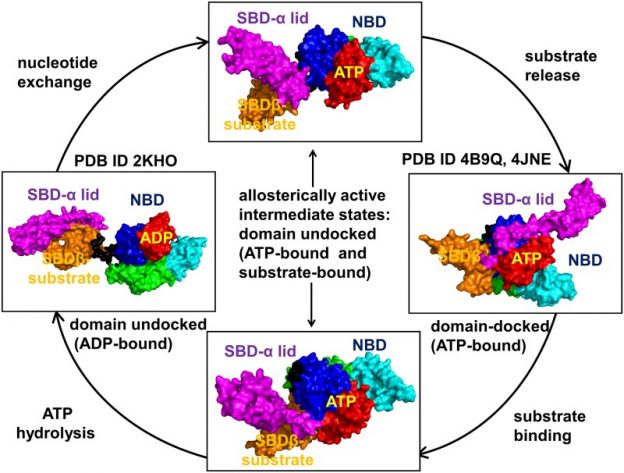
Dancing through Life: Molecular Dynamics Simulations and Network-Centric Modeling of Allosteric Mechanisms in Hsp70 and Hsp110 Chaperone Proteins.
Posted: September 12, 2017
Stetz G1, Verkhivker GM1,2. Author information 1 Graduate Program in Computational and Data Sciences, Schmid College of Science and Technology, Chapman University, Orange, California, United States of America. 2 Chapman […]
